Select image to enlarge
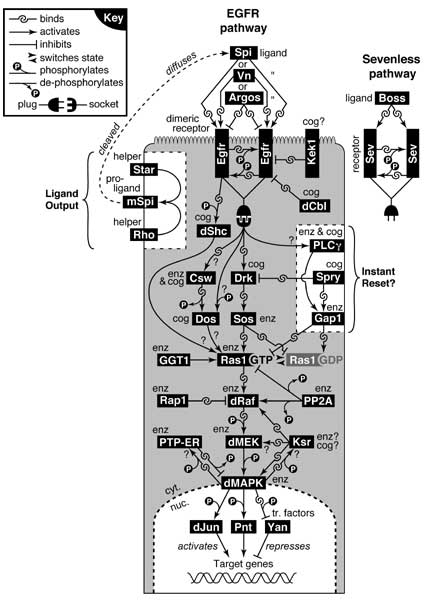
Figure 6.12
The EGFR and Sevenless signaling pathways.
See text for proteins, abbreviations, and a summary of how the core chain operates. One cell is sketched with microvilli at top, though ligand contact may take place elsewhere on the surface. Black rectangles = proteins; gray area = cytoplasm (cyt.); dashed line below = nuclear membrane. Arrows (activation) and blunt —| lines (repression) denote epistatic relations. Noncovalent binding is symbolized by interlocking hooks (cf. key at top). Question marks concern how or whether interactions occur. A 'cog' (a.k.a. 'adaptor') is a component that plays a steric (vs. enzymatic) role [441, 1954, 3299, 3676] (e.g., Drk [3185]), while 'enz' = enzyme. See also App. 7
Egfr is depicted at the apex but it actually affiliates with apico-lateral adherens junctions, as do Kek1 and dCbl [1001]. Conversely, the agents that process mSpi (white box at left) [199, 2244, 3385] are shown on the lateral membrane, but they may act apically [3692, 4192] via an ADAM-like protease [4558]. White box at right is a possible module for resetting Ras1 to an OFF state after signal transmission [3455]. Ras1 resides constitutively on the inner face of the membrane [781, 2653], as do Gap1 [3455], Csw [71], Dos [243], Sos [2137] (flies thus differ from mammals [106, 3485]), and Spry [681, 4340].
The Boss-Sev ligand-receptor complex is shown at right. It plugs into the same downstream circuitry as Egfr (sans Kek1, dShc, or the mSpi add-on) [1081, 3941], though effects of sevGOF and EgfrGOF can differ [1049]. The 7-pass transmembrane ligand Boss (Bride of sevenless) [1732] activates the RTK Sev (Sevenless) [1733, 2319, 2450, 3671, 4359]. Sev differs from Egfr in that its N-terminus is anchored to the membrane to form a big loop [226, 428, 1677] of unknown shape [3136]. Strangely, the entire Boss molecule (ripped from its own membrane?) is internalized by the Sev-expressing cell [601, 2318, 4211]. Sev is concentrated apically [194, 195] and at adherens junctions [4360]. Parochial players in the Sevenless (Sev) and Torso pathways (e.g., dCdc37 [932]) are omitted.
Binding partners in the core chain (listed from surface to DNA) are based on fly studies or vertebrate homologs (VH): Egfr-Drk [3829] (cf. Sev-Drk [3488]Δ), Drk-Sos [3488]Δ, Sos-Ras1 (VH [402, 4705]), Ras1GTP-dRaf (VH [2913]Δ), and dRaf-dMEK (VH [898, 2912]).
Ancillary binding partners (listed alphabetically, upstream factor first): Csw-Dos [243]Δ, dCbl-Spry (not shown) [4738], Egfr-Csw (probably indirect based on Sev-Csw [71]), Egfr-dShc (VH [4445]), Gap1-Ras1 (VH [3778, 4044]), Kek1-Egfr [1445], Ksr-dMAPK [2007]Δ, Ksr-dRaf [4285], dMAPK-PTP-ER [2133], dMAPK-Yan [2007], Rap1-dRaf (VH [3057]), Spry-Drk [681], and Spry-Gap1 [681]. Star apparently binds mSpi [1915].
This diagram is adapted from [2131, 3348, 3672, 3782]. It is augmented with data from sources cited above.
|
|
